library(car)
library(boot)
A <- runif(150)
B <- -2*A + runif(150)
C <- 3*B + runif(150)
x <- cbind(A, B, C)
par1 = 1 #column number of endogenous variable (Y)
par2 = 2 #column number of exogenous variable (X)
par3 = TRUE #use a constant term?
ylab = 'Y Variable Name'
xlab = 'X Variable Name'
main = 'Title Goes Here'
rsq <- function(formula, data, indices) {
d <- data[indices,] # allows boot to select sample
fit <- lm(formula, data=d)
return(summary(fit)$r.square)
}
cat("Selected columns\n")
(V1<-dimnames(x)[[2]][par1])
(V2<-dimnames(x)[[2]][par2])
cat("\n")
xdf <- data.frame(x[,par1], x[,par2])
names(xdf)<-c('Y', 'X')
if(par3 == FALSE) lmxdf<-lm(Y ~ X - 1, data = xdf) else lmxdf<-lm(Y~ X, data = xdf)
results <- boot(data=xdf, statistic=rsq, R=1000, formula=Y~X)
cat("\nResult of Simple Linear Regression computation\n")
(sumlmxdf<-summary(lmxdf))
cat("\nFitting an Analysis of Variance Model\n")
(aov.xdf<-aov(lmxdf) )
cat("\n\n\n")
(anova.xdf<-anova(lmxdf) )
cat("\n\n95% Confidence Interval of R-squared\n")
paste('[',round(boot.ci(results,type='bca')$bca[1,4], digits=3),',', round(boot.ci(results,type='bca')$bca[1,5], digits=3), ']',sep='')
plot(Y~ X, data=xdf, xlab=V2, ylab=V1, main='Regression Solution')
if(par3 == TRUE) abline(coef(lmxdf), col='red')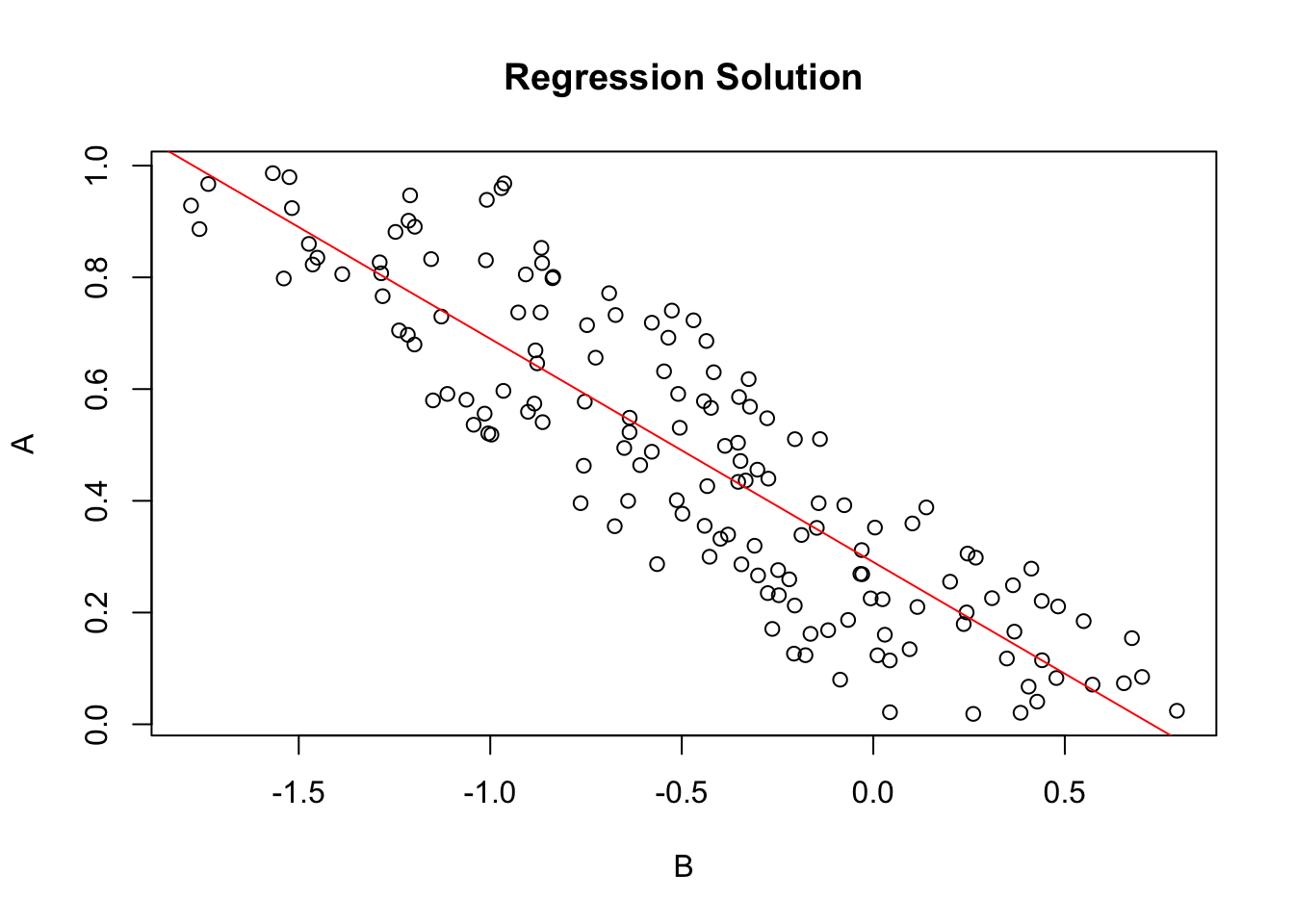
if(par3 == FALSE) abline(0.0, coef(lmxdf), col='red')
qqPlot(resid(lmxdf), main='QQplot of Residuals of Fit')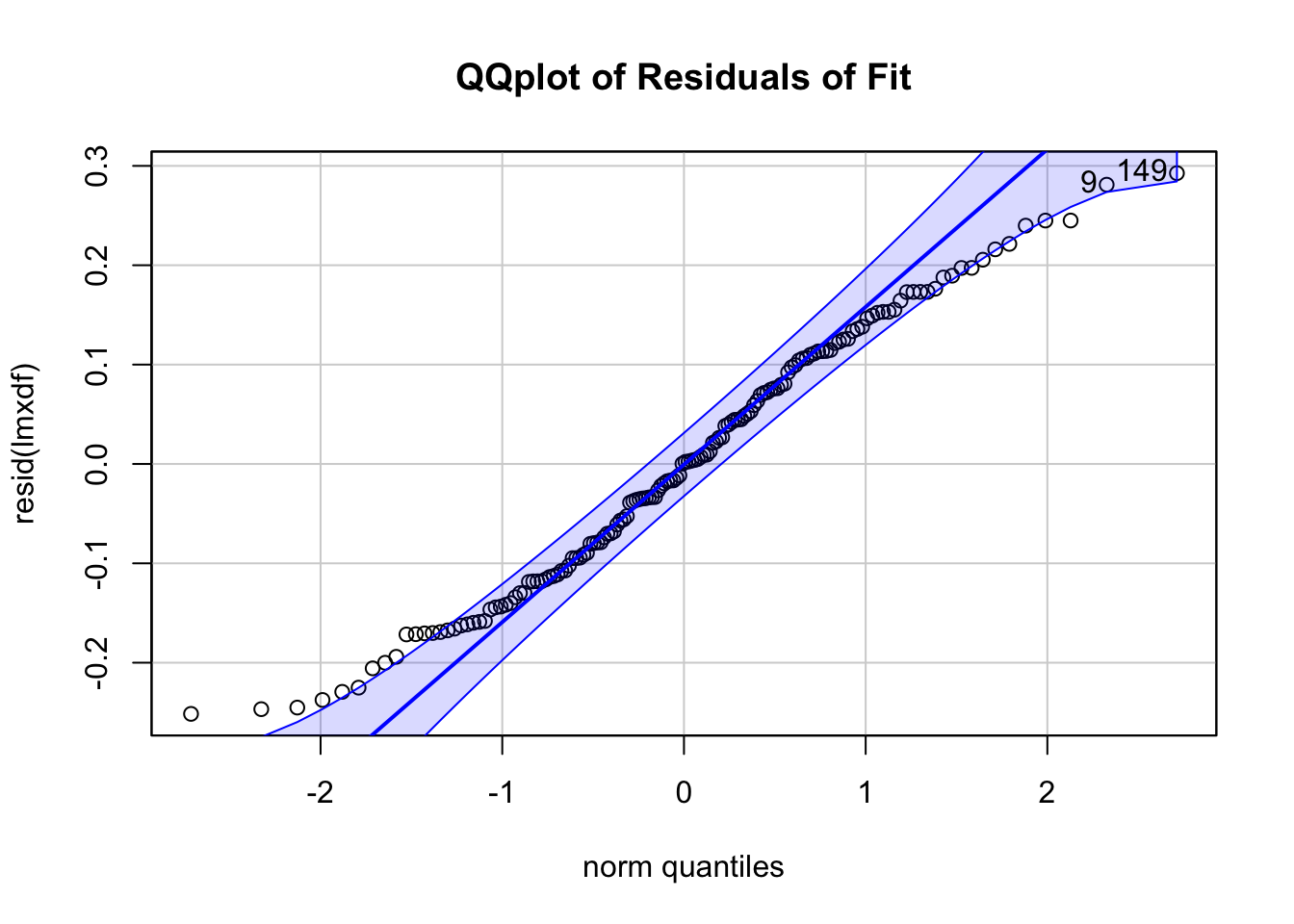
plot(xdf$X, resid(lmxdf), main='Scatterplot of Residuals of Model Fit')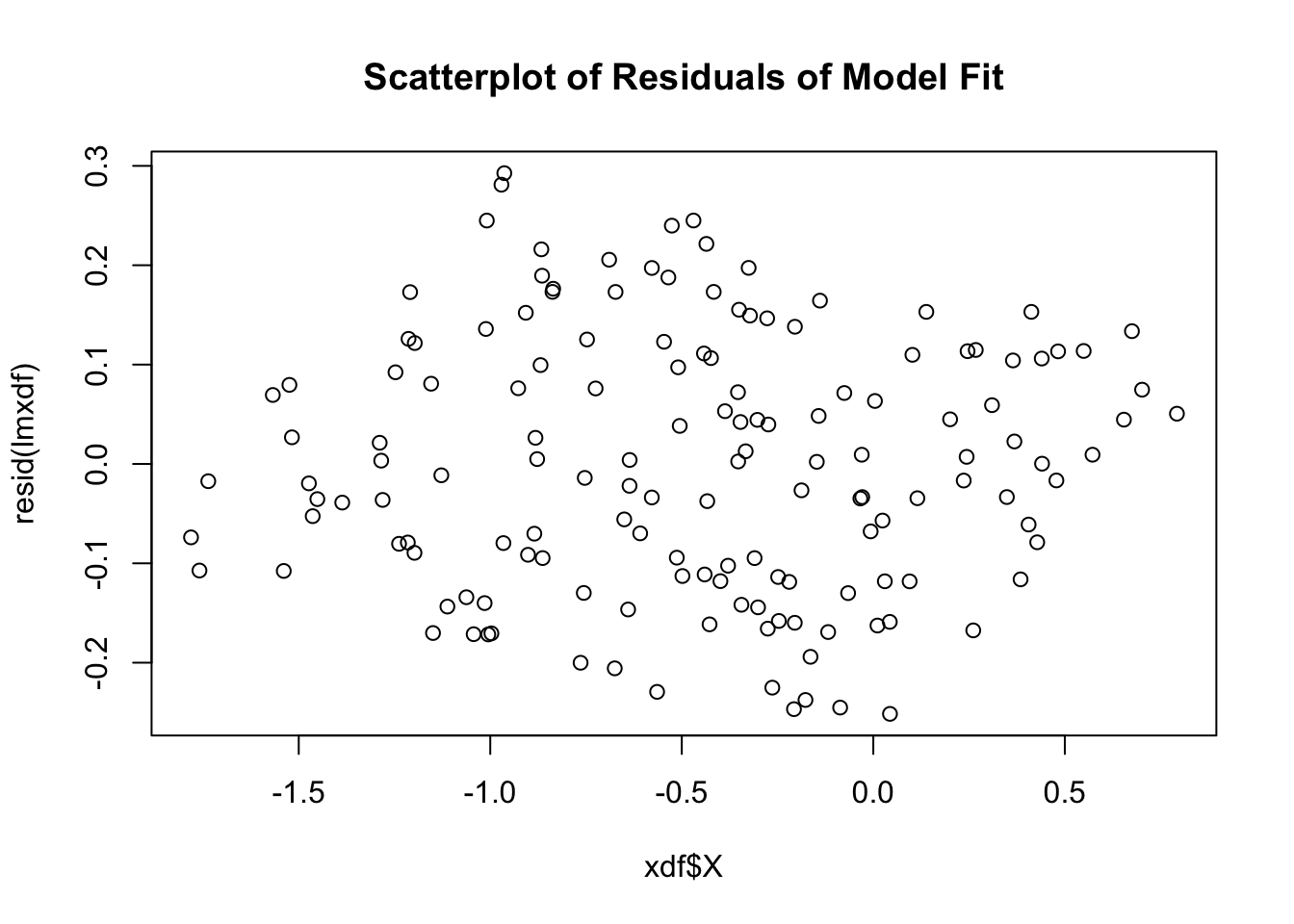
plot(lmxdf, which=4)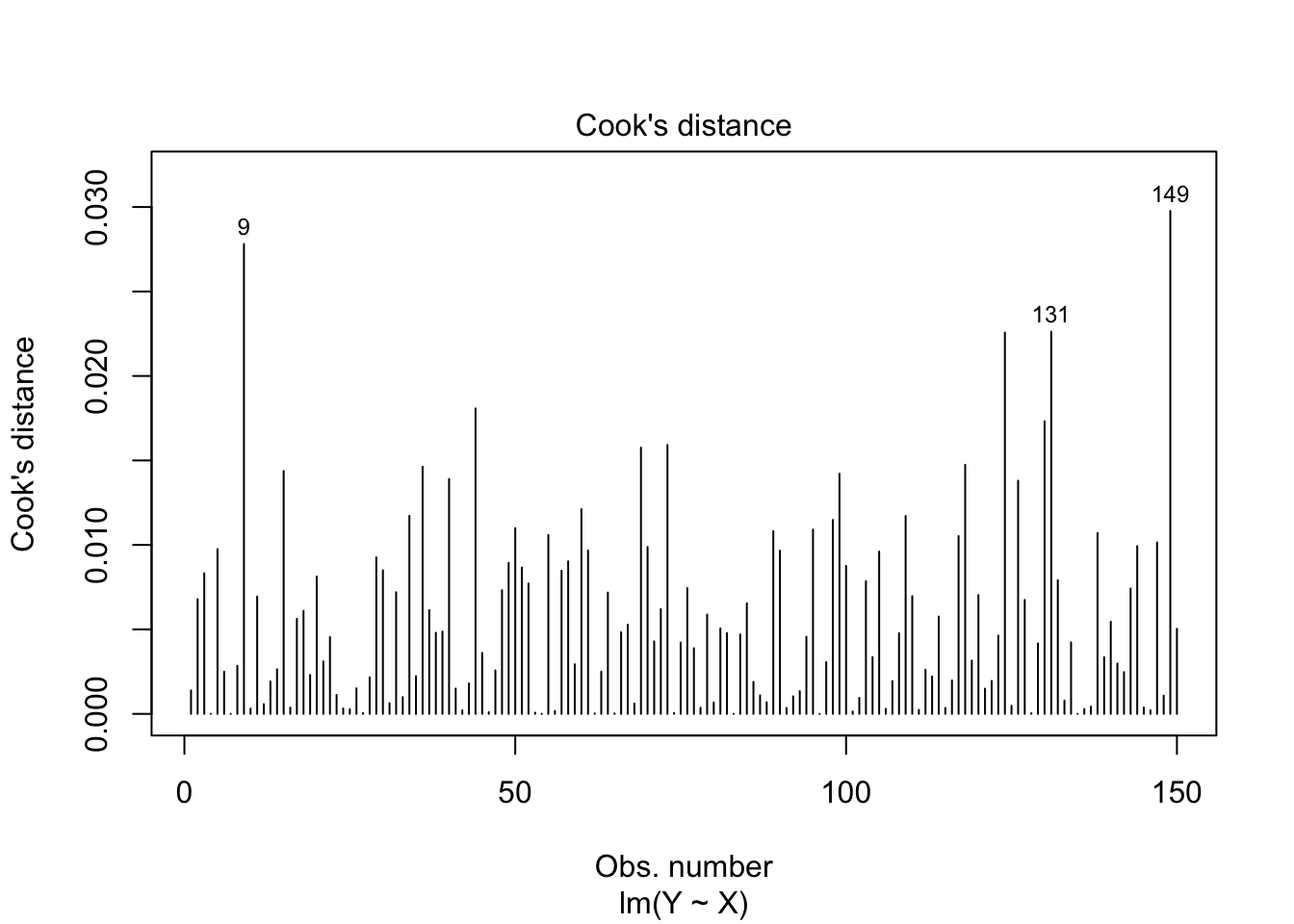
Selected columns
[1] "A"
[1] "B"
Result of Simple Linear Regression computation
Call:
lm(formula = Y ~ X, data = xdf)
Residuals:
Min 1Q Median 3Q Max
-0.251522 -0.107559 0.001211 0.106425 0.292664
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.29044 0.01349 21.53 <2e-16 ***
X -0.39972 0.01793 -22.30 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.1294 on 148 degrees of freedom
Multiple R-squared: 0.7706, Adjusted R-squared: 0.7691
F-statistic: 497.2 on 1 and 148 DF, p-value: < 2.2e-16
Fitting an Analysis of Variance Model
Call:
aov(formula = lmxdf)
Terms:
X Residuals
Sum of Squares 8.322257 2.477201
Deg. of Freedom 1 148
Residual standard error: 0.1293748
Estimated effects may be unbalanced
Analysis of Variance Table
Response: Y
Df Sum Sq Mean Sq F value Pr(>F)
X 1 8.3223 8.3223 497.21 < 2.2e-16 ***
Residuals 148 2.4772 0.0167
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
95% Confidence Interval of R-squared
[1] "[0.721,0.814]"
[1] 149 9